What is SCORE?
SCORE stands for Systematic CRISPR Outcome and Risk Evaluation, a machine-learning-based framework for modeling and predicting precision editing outcomes across genomic contexts.
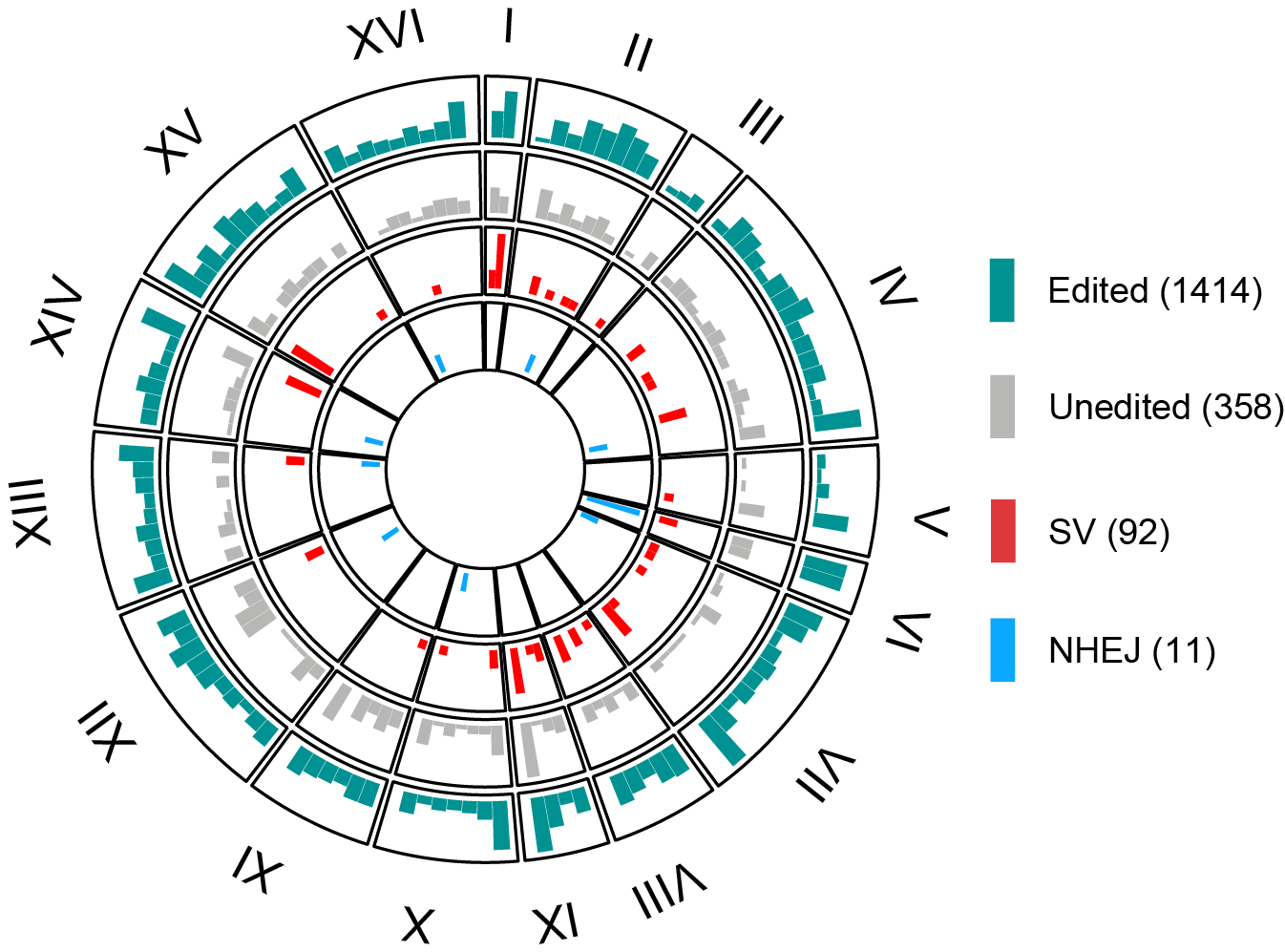
The training data of SCORE was a collection of 2,260 whole-genome sequencing datasets of Saccharomyces cerevisiae, wherein each single genome was treated with a single-guide RNA (sgRNA) for targeted cleavage of a unique genomic locus, paired with a double-stranded donor DNA template for the introduction of a natural SNP, indel or MNP genetic variant.
How does SCORE work?
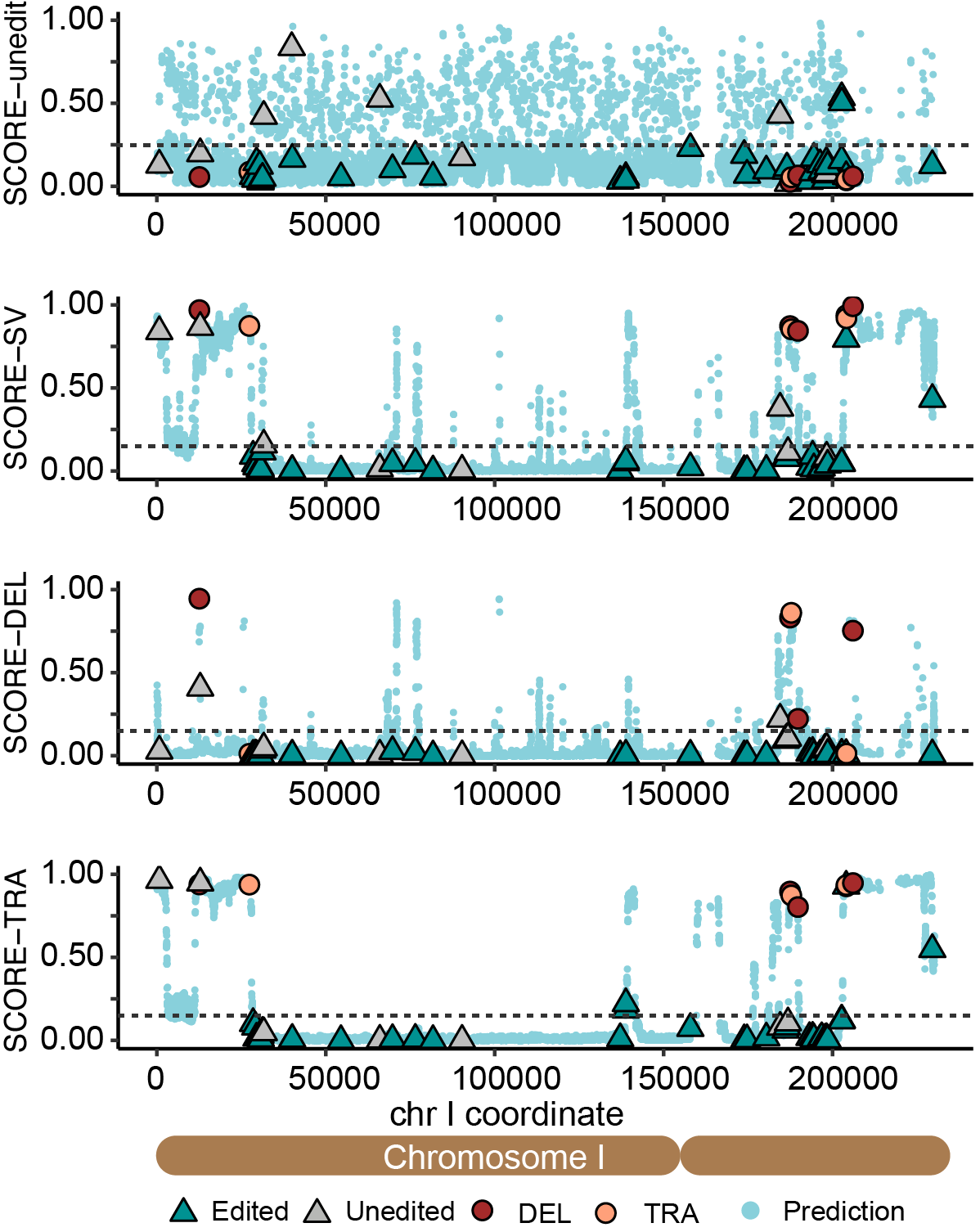
Through the training process, SCORE learnt the connection between genomic contexts of the cutting site (i.e. sequence composition, chromatin organization, sequence repetitiveness) and the consequence of editing (i.e. non-edits, errorneous edits). The resulted model is capable of predicting Cas9-induced structural alterations from the genomic contexts of the designed target site. Currently, SCORE contains four predictors trained separately for scoring the likelihood of different types of unwanted outcomes: unedit, structural variants (SVs), large deletions (DELs) and non-reciprocal translocations (TRAs).
How to use SCORE for my experimental design?
There are several opitions to use our pre-trained model to assist the design of CRISPR oligo(s):
Web-based tools:
The prediction scores of all 873,419 NGG PAM sites (excluding ones with exact zero-mismatch off-target sites) can be queried with Region browser or Query search , up to 5000 input queries (i.e. gRNA sequences) can be checked in one batch.
Download pre-calculated scores:
Through Downloads the pre-calculated table of predicted scores and genomic context measurements per NGG site on the reference genome (S288c) can be obtained.
Train new models:
The current version of SCORE was trained specifically with experimental data generated for Saccharomyces cerevisiae. It is expected that the genomic dependency of editing outcomes may vary in other species and experimental contexts (i.e. HDR efficiency). The original code for training the model can be found at GitHub: https://github.com/shli-embl/MAGESTIC-SCORE .
For any question and suggestion, please contact us via email: shengdi.li@embl.de .
Display CRISPR/Cas9 editing SCOREs on the genome
Compute editing SCOREs for gRNA sequences/library
Dataset 1. Download full table
Description: SCORE predictions and full annotation of 873,419 NGG PAM sites on S288C reference genome
Download .tsv
Dataset 2. Download validated sites
Description: List of NGG PAM sites with WGS validation outcomes
Download .tsv
Dataset 3. Download annotation files
Description: Processed chromatin, histone modification annotations used for PAM sites annotation and SCORE model training.
The files are used by our MAGESTIC-SCORE workflow ( https://github.com/shli-embl/MAGESTIC-SCORE ) to generate the predictive model of CRISPR/Cas9 editing outcomes in yeast.
Download .tar.gz
Dataset 4. Download track files for SGD genome browser
Description: SCORE predictions and bulk SV hotspots are available as BED files, loaded into the SGD genome browser here
*Notes: choose 'Wiggle XYPlot' for displaying a score distribution, and 'CanvasFeatures' for displaying hotspots.
Download .bed for SCORE-unedit
Download .bed for SCORE-SV
Download .bed for SCORE-DEL
Download .bed for SCORE-TRA
Download .bed for SV hotspots